For billions of years, life on Earth has relied on a specific set of 20 amino acids to build proteins. These molecular building blocks are the foundation of cellular function, strung together in complex sequences to create the machinery that keeps organisms alive. But is this “alphabet” of 20 essential? Or could life have evolved with a simpler, leaner chemistry?
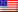
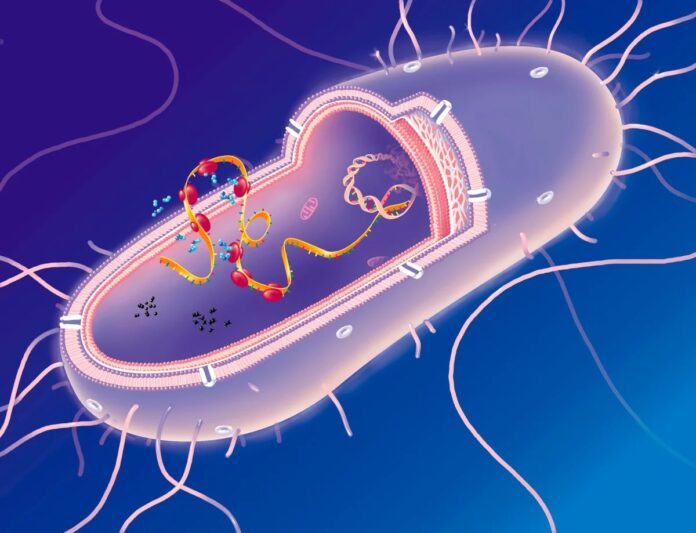
In a groundbreaking study published in Science, researchers from Columbia University, MIT, and Harvard University used artificial intelligence to engineer a strain of Escherichia coli bacteria capable of surviving with significantly reduced reliance on one of those 20 amino acids: isoleucine. While the bacteria still technically uses all 20 amino acids, the experiment demonstrates that core cellular machinery can be redesigned to function with far fewer components, hinting at how early life might have looked and showcasing the power of AI in synthetic biology.
Probing the Origins of Life
The primary goal of the research, led by Harris H. Wang, a professor of systems biology at Columbia University, was to explore the fundamental constraints of life’s chemistry. Scientists believe all modern life descends from a single-celled ancestor that lived over four billion years ago. However, theories suggest that even earlier, prebiotic life forms may have operated with a simpler chemical toolkit.
“Think about language. There are 26 letters in the English alphabet, but do you really need 26, or can you simplify that to 25 or 24?” says Wang. “The underlying question we seek to ask is what early life looks like.”
To test this, the team chose to target isoleucine, an amino acid structurally similar to valine and leucine. The hypothesis was that proteins might tolerate the removal of isoleucine if it were replaced by one of its close chemical relatives. They focused on the ribosome—the cell’s protein-building factory—which consists of more than 50 proteins. By redesigning these critical components, the researchers aimed to see if the cell could maintain high fitness levels despite this genetic simplification.
From Brute Force to AI Precision
The initial attempt to create this simplified organism relied on a “brute force” approach. The researchers identified 39 essential or highly expressed genes in E. coli and replaced every instance of isoleucine with valine or leucine. While the bacteria survived, their fitness dropped drastically to about 40% of normal wild-type levels. The team’s target was 90% fitness, indicating that simple substitution was not enough.
To bridge this gap, the researchers turned to artificial intelligence. They combined two powerful types of AI models:
- Sequence-based models (such as ESM2 and MSA Transformer) analyzed protein sequences to suggest evolutionarily plausible mutations that went beyond simple swaps.
- Structure-based models (such as AlphaFold2 and ProteinMPNN) verified that the redesigned proteins would fold into the correct three-dimensional shapes and interact properly with neighboring molecules.
The results were unexpected. The AI proposed complex structural changes that human researchers would not have anticipated. For example, when redesigning a ribosomal protein called RpsJ, the AI didn’t just swap amino acids; it remodeled an entire alpha helix and introduced eight new nearby mutations to compensate for the removal of just two isoleucines.
“Maybe these machine-learning systems know some aspects of biology we can experimentally verify but we don’t yet understand,” Wang notes.
Engineering a Robust Strain
Implementing these AI-designed changes was not straightforward. When tested individually, the mutations met the 90% fitness goal. However, when combined, they proved lethal to the cells. The team had to manually “debug” the genome, adding AI-designed segments in small batches to identify and fix lethal interactions.
The resulting strain, named Ec19, represents a significant milestone. It carries 21 ribosomal proteins that are completely free of isoleucine, alongside AI-redesigned versions of others. Crucially, Ec19 is robust:
* Its fitness remains above 90% of wild-type E. coli.
* The changes remained stable over 450 generations, showing no signs of reverting to the original genetic code under natural selection.
Implications for Biology and Beyond
While Ec19 is not a true 19-amino-acid organism—it still uses isoleucine in more than 81,000 other residues across its genome—it proves that life’s most ancient machinery can tolerate substantial simplification. This work highlights the evolving role of AI in synthetic biology.
Tom Ellis, a professor of synthetic genome engineering at Imperial College London, described the study as a “tour de force.” He noted that the rapid advancement of AI-enabled protein modeling over the last seven years has made such complex engineering possible.
The implications extend beyond understanding Earth’s history. This technology could inform biotechnology applications in extreme environments, such as space exploration, where not every amino acid may be readily available. As DNA synthesis becomes cheaper and AI models become more capable of handling whole genomes, scientists may one day create organisms that truly run on a reduced chemical alphabet, fundamentally altering our understanding of life’s possibilities.